Challenge Data¶
We gratefully acknowledge our sponsor, Climb 4 Kidney Cancer (C4KC), for their generous support which made the collection and annotation of this data possible.
Access the Data¶
The 210 patients of training data were made available on GitHub on March 15, 2019. The imaging alone for the remaining 90 patients will be made available on July 15, 2019, two weeks prior to the July 29 deadline for submitting test predictions. All data is licensed as CC BY-NC-SA.
Please note that we consider benchmarking for the purpose of publication on MLCommons to be acceptable use under this license.
Data Description¶
This is an abbreviated version of the complete data description manuscript.
Cohort¶
All patients who underwent partial or radical nephrectomy for one or more kidney tumors at the University of Minnesota Medical Center between 2010 and 2018 were candidates for inclusion in this database. Cases where we could not find a pre-operative arterial phase abdominal CT were excluded. From the remaining cases, 300 were selected at random.
Data Format¶
The imaging as well as ground truth labels were provided in an anonymized NIFTI format with shape (num_slices, height, width). Here, num_slices correspond to an axial view, and progress from superior to inferior as the slice index increases. In all cases, the patient was supine during image collection, and the height-width origins thus lie to the patients' left anteriors. When there were multiple qualifying series for a particular case, that with the smaller slice thickness was chosen, but slice thicknesses range from 1mm to 5mm.
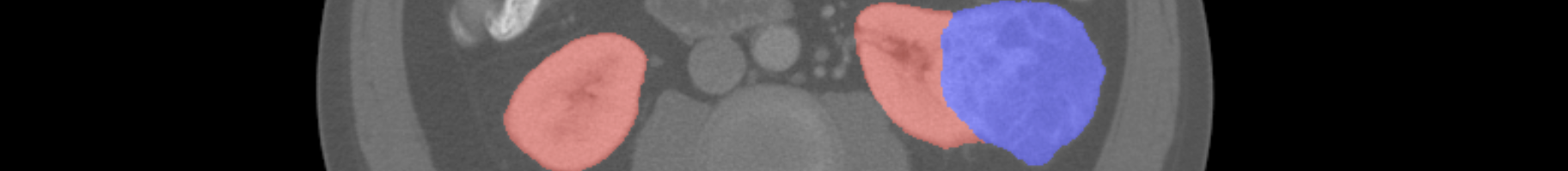
Ground Truth Labels¶
Manual segmentation labels were provided by medical students under the supervision of our clinical chair, Dr. Christopher Weight. The annotation procedure was as follows:
- The first and last slices containing kidney or tumor tissue were recorded. We refer to them as pre and post, respectively.
- The first and last slices containing a renal hilum for each kidney were recorded. We refer to them as lhb, lhe, rhb, and rhe for left hilum begin, left hilum end, right hilum begin, and right hilum end respectively.
- Evenly spaced slices were selected for annotation (as few as 1 out of every 5) such that the largest number of selected slices between pre and post was below 50 if possible.
- For each selected slice, the annotator drew one or more contours such that all tissue depicted inside the contours was kidney, tumor, or fat. We refer to these as kidney contours.
- In slices where the renal hilum was present, the annotator excluded some of the sinus and collecting system from their contours such that the largest concavity in their contour began and ended at the entrance to the hilum.
- The annotator then drew a new series of contours such that the only tissue depicted in their interiors was tumor or not kidney. We refer to these as tumor contours.
The final ground truth label was computed from these annotations as follows:
- Kidney contours for unannotated slices were interpolated from their nearest annotated slices on either side. This was done by simple linear morphing.
- For each slice between pre and post, a simple 3x3 Gaussian blur was performed, and pixels inside kidney contours below a blurred Hounsfield Unit value of -30 were excluded as likely fat [1].
- For each contour containing a hilum, a line was drawn such that the largest deficiency in convexity was filled, and pixels within this deficiency above the above HU threshold were included.
- Tumor contours were also interpolated with a simple morph.
- For each tumor contour, a pixel-wise AND was performed between its interior pixels and the thresholded kidney. Remaining pixels are considered to be tumor.
This procedure worked for a vast majority of cases, but occasionally there were other concavities that were larger than that of the hilum. In these cases, the endpoints of the hilum lines were chosen manually
Annotators had access to each case's attending radiologist's conclusion, as well as the conclusion from surgical pathology, which helped to facilitate accurate localization of tumors and exclusion of cysts. All cases were then reviewed in both the axial and coronal planes. Corrections were made by Nicholas Heller under the direction of Drs. Christopher Weight and Niranjan Sathianathen where necessary.
Below is a list of students who contributed to this effort:
- Heather Kaluzniak
- Keenan Moore
- Joel Rosenberg
- Paul Blake
- Makinna Oestreich
- Ed Walczak
- Zach Rengel
- Shane Raza
- Zach Edgerton
- Michael Tradewell
- Aneri Shah
References¶
1. Page 83 in: Herbert Lepor (2000). Prostatic Diseases. W.B. Saunders Company. ISBN 9780721674162.